Human Cyclin-dependent kinase 5 (CDK5) ELISA Kit (HUEB0834)
- SKU:
- HUEB0834
- Product Type:
- ELISA Kit
- Size:
- 96 Assays
- Uniprot:
- Q00535
- Range:
- 0.312-20 ng/mL
- ELISA Type:
- Sandwich
- Synonyms:
- CDK5, Cyclin-dependent kinase 5, TPKII catalytic subunit, Tau protein kinase II catalytic subunit, Cell division protein kinase 5, Serine, threonine-protein kinase PSSALRE
- Reactivity:
- Human
Description
Human Cyclin-dependent kinase 5 (CDK5) ELISA Kit
The Human Cyclin-Dependent Kinase 5 (CDK5) ELISA Kit is specifically designed for the precise measurement of CDK5 levels in human samples including serum, plasma, and cell culture supernatants. This kit offers exceptional sensitivity and specificity, ensuring accurate and consistent results for various research applications.CDK5 is a key enzyme that plays a critical role in cell cycle regulation, neuronal development, and synaptic plasticity. Dysregulation of CDK5 has been linked to neurodegenerative diseases such as Alzheimer's and Parkinson's, as well as various cancers.
As a result, CDK5 serves as a vital biomarker for studying these conditions and investigating potential therapeutic interventions.With the Human CDK5 ELISA Kit, researchers can gain valuable insights into the mechanisms underlying CDK5-related diseases and facilitate the development of novel treatment strategies. Trust in the reliability and precision of this kit for your research needs in the field of cell biology and neurology.
| Product Name: | Human Cyclin-dependent kinase 5 (CDK5) ELISA Kit |
| SKU: | HUEB0834 |
| Size: | 96T |
| Target: | Human Cyclin-dependent kinase 5 (CDK5) |
| Synonyms: | Cell division protein kinase 5, Serine/threonine-protein kinase PSSALRE, Tau protein kinase II catalytic subunit, TPKII catalytic subunit, CDKN5 |
| Assay Type: | Sandwich |
| Detection Method: | ELISA |
| Reactivity: | Human |
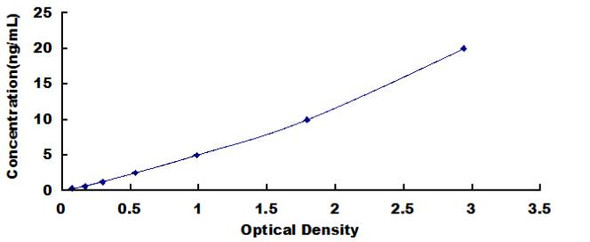
| Detection Range: | 0.312-20ng/mL |
| Sensitivity: | 0.1ng/mL |
| Intra CV: | 6.1% | ||||||||||||||||||||
| Inter CV: | 9.3% | ||||||||||||||||||||
| Linearity: |
| ||||||||||||||||||||
| Recovery: |
| ||||||||||||||||||||
| Function: | Proline-directed serine/threonine-protein kinase essential for neuronal cell cycle arrest and differentiation and may be involved in apoptotic cell death in neuronal diseases by triggering abortive cell cycle re-entry. Interacts with D1 and D3-type G1 cyclins. Phosphorylates SRC, NOS3, VIM/vimentin, p35/CDK5R1, MEF2A, SIPA1L1, SH3GLB1, PXN, PAK1, MCAM/MUC18, SEPT5, SYN1, DNM1, AMPH, SYNJ1, CDK16, RAC1, RHOA, CDC42, TONEBP/NFAT5, MAPT/TAU, MAP1B, histone H1, p53/TP53, HDAC1, APEX1, PTK2/FAK1, huntingtin/HTT, ATM, MAP2, NEFH and NEFM. Regulates several neuronal development and physiological processes including neuronal survival, migration and differentiation, axonal and neurite growth, synaptogenesis, oligodendrocyte differentiation, synaptic plasticity and neurotransmission, by phosphorylating key proteins. Activated by interaction with CDK5R1 (p35) and CDK5R2 (p39), especially in post-mitotic neurons, and promotes CDK5R1 (p35) expression in an autostimulation loop. Phosphorylates many downstream substrates such as Rho and Ras family small GTPases (e.g. PAK1, RAC1, RHOA, CDC42) or microtubule-binding proteins (e.g. MAPT/TAU, MAP2, MAP1B), and modulates actin dynamics to regulate neurite growth and/or spine morphogenesis. Phosphorylates also exocytosis associated proteins such as MCAM/MUC18, SEPT5, SYN1, and CDK16/PCTAIRE1 as well as endocytosis associated proteins such as DNM1, AMPH and SYNJ1 at synaptic terminals. In the mature central nervous system (CNS), regulates neurotransmitter movements by phosphorylating substrates associated with neurotransmitter release and synapse plasticity; synaptic vesicle exocytosis, vesicles fusion with the presynaptic membrane, and endocytosis. Promotes cell survival by activating anti-apoptotic proteins BCL2 and STAT3, and negatively regulating of JNK3/MAPK10 activity. Phosphorylation of p53/TP53 in response to genotoxic and oxidative stresses enhances its stabilization by preventing ubiquitin ligase-mediated proteasomal degradation, and induces transactivation of p53/TP53 target genes, thus regulating apoptosis. Phosphorylation of p35/CDK5R1 enhances its stabilization by preventing calpain-mediated proteolysis producing p25/CDK5R1 and avoiding ubiquitin ligase-mediated proteasomal degradation. During aberrant cell-cycle activity and DNA damage, p25/CDK5 activity elicits cell-cycle activity and double-strand DNA breaks that precedes neuronal death by deregulating HDAC1. DNA damage triggered phosphorylation of huntingtin/HTT in nuclei of neurons protects neurons against polyglutamine expansion as well as DNA damage mediated toxicity. Phosphorylation of PXN reduces its interaction with PTK2/FAK1 in matrix-cell focal adhesions (MCFA) during oligodendrocytes (OLs) differentiation. Negative regulator of Wnt/beta-catenin signaling pathway. Activator of the GAIT (IFN-gamma-activated inhibitor of translation) pathway, which suppresses expression of a post-transcriptional regulon of proinflammatory genes in myeloid cells; phosphorylates the linker domain of glutamyl-prolyl tRNA synthetase (EPRS) in a IFN-gamma-dependent manner, the initial event in assembly of the GAIT complex. Phosphorylation of SH3GLB1 is required for autophagy induction in starved neurons. Phosphorylation of TONEBP/NFAT5 in response to osmotic stress mediates its rapid nuclear localization. MEF2 is inactivated by phosphorylation in nucleus in response to neurotoxin, thus leading to neuronal apoptosis. APEX1 AP-endodeoxyribonuclease is repressed by phosphorylation, resulting in accumulation of DNA damage and contributing to neuronal death. NOS3 phosphorylation down regulates NOS3-derived nitrite (NO) levels. SRC phosphorylation mediates its ubiquitin-dependent degradation and thus leads to cytoskeletal reorganization. May regulate endothelial cell migration and angiogenesis via the modulation of lamellipodia formation. Involved in dendritic spine morphogenesis by mediating the EFNA1-EPHA4 signaling. The complex p35/CDK5 participates in the regulation of the circadian clock by modulating the function of CLOCK protein: phosphorylates CLOCK at 'Thr-451' and 'Thr-461' and regulates the transcriptional activity of the CLOCK-ARNTL/BMAL1 heterodimer in association with altered stability and subcellular distribution. |
| Uniprot: | Q00535 |
| Sample Type: | Serum, plasma, tissue homogenates, cell culture supernates and other biological fluids |
| Specificity: | Natural and recombinant human Cyclin-dependent-like kinase 5 |
| Sub Unit: | Heterodimer composed of a catalytic subunit CDK5 and a regulatory subunit CDK5R1 (p25) and macromolecular complex composed of at least CDK5, CDK5R1 (p35) and CDK5RAP1 or CDK5RAP2 or CDK5RAP3. Only the heterodimer shows kinase activity. Under neurotoxic stress and neuronal injury conditions, p35 is cleaved by calpain to generate p25 that hyperactivates CDK5, that becomes functionally disabled and often toxic. Found in a trimolecular complex with CABLES1 and ABL1. Interacts with CABLES1 and CABLES2 (By similarity). Interacts with AATK and GSTP1. Binds to HDAC1 when in complex with p25. Interaction with myristoylation p35 promotes CDK5 association with membranes. Both isoforms 1 and 2 interacts with beta-catenin/CTNNB1. Interacts with delta-catenin/CTNND2 and APEX1. Interacts with P53/TP53 in neurons. Interacts with EPHA4; may mediate the activation of NGEF by EPHA4. Interacts with PTK2/FAK1 (By similarity). The complex p35/CDK5 interacts with CLOCK. Interacts with HTR6. |
| Research Area: | Neurosciences |
| Subcellular Location: | Isoform 2 Nucleus |
| Storage: | Please see kit components below for exact storage details |
| Note: | For research use only |
| UniProt Protein Function: | CDK5: a protein kinase of the CDK family. Unlike other members of this family, it is not activated by cyclins but by p35 (CDK5R1) and p39. An important regulator of neuronal positioning during brain development. May also play a role in synaptogenesis and neurotransmission. Substrates include TAU, MAP2, NF-H and -M, Nudel, PDE6, beta-catenin, amphyphysin, dynamin I, synapsin 1, Munc-18, and NMDA receptor 2A. Plays a role in myogenesis, haematopoietic cell differentiation, spermatogenesis, insulin secretion, and lens differentiation. Implicated in the pathology of neurofibrillary tangles and formation of senile plaques, hallmarks of Alzheimer?s disease. Induces tau phosphorylation and aggregation and neurofibrillary tangle deposition and neurodegeneration in in vitro and in vivo animal models. Brain samples from Alzeimer?s pateints show elevated CDK5 activity. |
| UniProt Protein Details: | Protein type:Protein kinase, CMGC; Kinase, protein; Cell cycle regulation; Protein kinase, Ser/Thr (non-receptor); EC 2.7.11.1; CMGC group; CDK family; CDK5 subfamily; CDK/CDK5 subfamily Chromosomal Location of Human Ortholog: 7q36 Cellular Component: axon; cell junction; cell soma; cytoplasm; cytosol; dendrite; growth cone; membrane; neuromuscular junction; nucleoplasm; nucleus; plasma membrane; postsynaptic density Molecular Function:acetylcholine receptor activator activity; cyclin-dependent protein kinase activity; ErbB-2 class receptor binding; ErbB-3 class receptor binding; kinase activity; protein binding; protein kinase activity; protein serine/threonine kinase activity; tau-protein kinase activity Biological Process: axon extension; cell proliferation; negative regulation of proteolysis; negative regulation of transcription, DNA-dependent; neurite development; neuron apoptosis; neuron differentiation; neuron migration; oligodendrocyte differentiation; peptidyl-serine phosphorylation; phosphorylation; positive regulation of neuron apoptosis; regulation of apoptosis; regulation of macroautophagy; regulation of synaptic plasticity; synaptic transmission; synaptic vesicle endocytosis; synaptic vesicle exocytosis; synaptogenesis Disease: Lissencephaly 7 With Cerebellar Hypoplasia |
| NCBI Summary: | This gene encodes a proline-directed serine/threonine kinase that is a member of the cyclin-dependent kinase family of proteins. Unlike other members of the family, the protein encoded by this gene does not directly control cell cycle regulation. Instead the protein, which is predominantly expressed at high levels in mammalian postmitotic central nervous system neurons, functions in diverse processes such as synaptic plasticity and neuronal migration through phosphorylation of proteins required for cytoskeletal organization, endocytosis and exocytosis, and apoptosis. In humans, an allelic variant of the gene that results in undetectable levels of the protein has been associated with lethal autosomal recessive lissencephaly-7. Alternative splicing results in multiple transcript variants. [provided by RefSeq, May 2015] |
| UniProt Code: | Q00535 |
| NCBI GenInfo Identifier: | 4033704 |
| NCBI Gene ID: | 1020 |
| NCBI Accession: | Q00535.3 |
| UniProt Secondary Accession: | Q00535,A1XKG3, |
| UniProt Related Accession: | Q00535 |
| Molecular Weight: | 29,544 Da |
| NCBI Full Name: | Cyclin-dependent-like kinase 5 |
| NCBI Synonym Full Names: | cyclin dependent kinase 5 |
| NCBI Official Symbol: | CDK5 |
| NCBI Official Synonym Symbols: | LIS7; PSSALRE |
| NCBI Protein Information: | cyclin-dependent-like kinase 5 |
| UniProt Protein Name: | Cyclin-dependent-like kinase 5 |
| UniProt Synonym Protein Names: | Cell division protein kinase 5; Serine/threonine-protein kinase PSSALRE; Tau protein kinase II catalytic subunit; TPKII catalytic subunit |
| Protein Family: | CDK5RAP1-like protein |
| UniProt Gene Name: | CDK5 |
| UniProt Entry Name: | CDK5_HUMAN |
| Component | Quantity (96 Assays) | Storage |
| ELISA Microplate (Dismountable) | 8×12 strips | -20°C |
| Lyophilized Standard | 2 | -20°C |
| Sample Diluent | 20ml | -20°C |
| Assay Diluent A | 10mL | -20°C |
| Assay Diluent B | 10mL | -20°C |
| Detection Reagent A | 120µL | -20°C |
| Detection Reagent B | 120µL | -20°C |
| Wash Buffer | 30mL | 4°C |
| Substrate | 10mL | 4°C |
| Stop Solution | 10mL | 4°C |
| Plate Sealer | 5 | - |
Other materials and equipment required:
- Microplate reader with 450 nm wavelength filter
- Multichannel Pipette, Pipette, microcentrifuge tubes and disposable pipette tips
- Incubator
- Deionized or distilled water
- Absorbent paper
- Buffer resevoir
*Note: The below protocol is a sample protocol. Protocols are specific to each batch/lot. For the correct instructions please follow the protocol included in your kit.
Allow all reagents to reach room temperature (Please do not dissolve the reagents at 37°C directly). All the reagents should be mixed thoroughly by gently swirling before pipetting. Avoid foaming. Keep appropriate numbers of strips for 1 experiment and remove extra strips from microtiter plate. Removed strips should be resealed and stored at -20°C until the kits expiry date. Prepare all reagents, working standards and samples as directed in the previous sections. Please predict the concentration before assaying. If values for these are not within the range of the standard curve, users must determine the optimal sample dilutions for their experiments. We recommend running all samples in duplicate.
| Step | |
| 1. | Add Sample: Add 100µL of Standard, Blank, or Sample per well. The blank well is added with Sample diluent. Solutions are added to the bottom of micro ELISA plate well, avoid inside wall touching and foaming as possible. Mix it gently. Cover the plate with sealer we provided. Incubate for 120 minutes at 37°C. |
| 2. | Remove the liquid from each well, don't wash. Add 100µL of Detection Reagent A working solution to each well. Cover with the Plate sealer. Gently tap the plate to ensure thorough mixing. Incubate for 1 hour at 37°C. Note: if Detection Reagent A appears cloudy warm to room temperature until solution is uniform. |
| 3. | Aspirate each well and wash, repeating the process three times. Wash by filling each well with Wash Buffer (approximately 400µL) (a squirt bottle, multi-channel pipette,manifold dispenser or automated washer are needed). Complete removal of liquid at each step is essential. After the last wash, completely remove remaining Wash Buffer by aspirating or decanting. Invert the plate and pat it against thick clean absorbent paper. |
| 4. | Add 100µL of Detection Reagent B working solution to each well. Cover with the Plate sealer. Incubate for 60 minutes at 37°C. |
| 5. | Repeat the wash process for five times as conducted in step 3. |
| 6. | Add 90µL of Substrate Solution to each well. Cover with a new Plate sealer and incubate for 10-20 minutes at 37°C. Protect the plate from light. The reaction time can be shortened or extended according to the actual color change, but this should not exceed more than 30 minutes. When apparent gradient appears in standard wells, user should terminatethe reaction. |
| 7. | Add 50µL of Stop Solution to each well. If color change does not appear uniform, gently tap the plate to ensure thorough mixing. |
| 8. | Determine the optical density (OD value) of each well at once, using a micro-plate reader set to 450 nm. User should open the micro-plate reader in advance, preheat the instrument, and set the testing parameters. |
| 9. | After experiment, store all reagents according to the specified storage temperature respectively until their expiry. |
When carrying out an ELISA assay it is important to prepare your samples in order to achieve the best possible results. Below we have a list of procedures for the preparation of samples for different sample types.
| Sample Type | Protocol |
| Serum | If using serum separator tubes, allow samples to clot for 30 minutes at room temperature. Centrifuge for 10 minutes at 1,000x g. Collect the serum fraction and assay promptly or aliquot and store the samples at -80°C. Avoid multiple freeze-thaw cycles. If serum separator tubes are not being used, allow samples to clot overnight at 2-8°C. Centrifuge for 10 minutes at 1,000x g. Remove serum and assay promptly or aliquot and store the samples at -80°C. Avoid multiple freeze-thaw cycles. |
| Plasma | Collect plasma using EDTA or heparin as an anticoagulant. Centrifuge samples at 4°C for 15 mins at 1000 × g within 30 mins of collection. Collect the plasma fraction and assay promptly or aliquot and store the samples at -80°C. Avoid multiple freeze-thaw cycles. Note: Over haemolysed samples are not suitable for use with this kit. |
| Urine & Cerebrospinal Fluid | Collect the urine (mid-stream) in a sterile container, centrifuge for 20 mins at 2000-3000 rpm. Remove supernatant and assay immediately. If any precipitation is detected, repeat the centrifugation step. A similar protocol can be used for cerebrospinal fluid. |
| Cell culture supernatant | Collect the cell culture media by pipette, followed by centrifugation at 4°C for 20 mins at 1500 rpm. Collect the clear supernatant and assay immediately. |
| Cell lysates | Solubilize cells in lysis buffer and allow to sit on ice for 30 minutes. Centrifuge tubes at 14,000 x g for 5 minutes to remove insoluble material. Aliquot the supernatant into a new tube and discard the remaining whole cell extract. Quantify total protein concentration using a total protein assay. Assay immediately or aliquot and store at ≤ -20 °C. |
| Tissue homogenates | The preparation of tissue homogenates will vary depending upon tissue type. Rinse tissue with 1X PBS to remove excess blood & homogenize in 20ml of 1X PBS (including protease inhibitors) and store overnight at ≤ -20°C. Two freeze-thaw cycles are required to break the cell membranes. To further disrupt the cell membranes you can sonicate the samples. Centrifuge homogenates for 5 mins at 5000xg. Remove the supernatant and assay immediately or aliquot and store at -20°C or -80°C. |
| Tissue lysates | Rinse tissue with PBS, cut into 1-2 mm pieces, and homogenize with a tissue homogenizer in PBS. Add an equal volume of RIPA buffer containing protease inhibitors and lyse tissues at room temperature for 30 minutes with gentle agitation. Centrifuge to remove debris. Quantify total protein concentration using a total protein assay. Assay immediately or aliquot and store at ≤ -20 °C. |
| Breast Milk | Collect milk samples and centrifuge at 10,000 x g for 60 min at 4°C. Aliquot the supernatant and assay. For long term use, store samples at -80°C. Minimize freeze/thaw cycles. |